```{r}
#| include: false
suppressPackageStartupMessages(library(tidyverse))
suppressPackageStartupMessages(library(knitr))
suppressPackageStartupMessages(library(htmltools))
source(normalizePath(file.path(dirname(knitr::current_input(dir = TRUE)), "..", "R", "chapter_table_helpers.R"), mustWork = TRUE))
source(normalizePath(file.path(dirname(knitr::current_input(dir = TRUE)), "..", "R", "chapter_helpers.R"), mustWork = TRUE))
```
:::: {.content-visible when-format="html"}
```{webr-r}
#| context: setup
#| include: false
#| echo: false
library(dplyr)
library(ggplot2)
library(tibble)
library(boot)
chapter10_format_p <- function(p, digits = 3) {
threshold <- 10^(-digits)
ifelse(
p < threshold,
paste0("< ", formatC(threshold, format = "f", digits = digits)),
formatC(p, format = "f", digits = digits)
)
}
```
::::
# Chapter 10: Exact Tests and Resampling Methods
Small-sample analysis often requires reference distributions that do not depend on large-sample theory.
Exact tests and resampling methods become especially useful when sample sizes are modest and large-sample approximations carry meaningful risk of inaccuracy. This chapter explains when to use exact tests for discrete outcomes, when permutation tests provide a cleaner reference distribution than a parametric model, and when bootstrap resampling is useful for interval estimation. The emphasis is practical: matching the method to the design, the outcome type, and the inferential goal.
### Learning Objectives
By the end of this chapter, you will be able to explain when exact tests are preferable to large-sample approximations, distinguish conditional exact tests from unconditional and mid-p alternatives, implement exact binomial, exact Poisson, permutation, and bootstrap procedures in R, and report resampling analyses with enough detail for readers to reproduce the statistic, number of resamples, random seed, and inferential target.
### When to Use Exact and Resampling Methods
Exact tests calculate p-values directly from the null distribution of the statistic. They are especially useful when the outcome is discrete, when expected cell counts are small, or when the sample is too limited for asymptotic results to be reliable.
Resampling methods use the observed data to approximate a sampling distribution. Permutation tests reassign labels under the null hypothesis to create a reference distribution for a test statistic. Bootstrap methods resample with replacement from the observed data to approximate the variability of an estimator and to construct confidence intervals.
In practice, exact tests are most natural for sparse binary or count data. Permutation tests are useful when exchangeability under the null is plausible. Bootstrap methods are especially helpful when the statistic of interest lacks a simple closed-form standard error.
For any resampling analysis, report the statistic being resampled and the number of permutations or bootstrap resamples. Also report the random seed and whether the result is exact or Monte Carlo. Those details determine the stability and reproducibility of the reported inference.
### Fisher's Exact Test for 2×2 Tables
Fisher's exact test is the standard conditional test for association in a 2×2 contingency table when some expected cell counts are small. It conditions on the observed margins and calculates the probability of the observed table, and all equally or more extreme tables, under the null hypothesis of no association.
Because the sample space is discrete, Fisher's test can be conservative: its p-value is guaranteed to control the Type I error rate, but that guarantee can come at the cost of reduced power. That trade-off is often acceptable in confirmatory work, especially when one of the margins is fixed by design.
```{r}
#| include: false
training_table <- matrix(
c(8, 2, 3, 7),
nrow = 2,
byrow = TRUE,
dimnames = list(
Training = c("Yes", "No"),
Target = c("Met", "Not Met")
)
)
training_display_table <- tibble(
`Training group` = c("Training", "No training"),
`Met target` = c(8, 3),
`Did not meet target` = c(2, 7)
)
training_expected_counts <- suppressWarnings(chisq.test(training_table)$expected)
training_min_expected <- min(training_expected_counts)
fisher_result <- fisher.test(training_table)
training_mosaic_data <- tibble(
`Training group` = c("Training", "Training", "No training", "No training"),
Target = c("Met", "Not Met", "Met", "Not Met"),
count = c(8, 2, 3, 7)
) %>%
mutate(
`Training group` = factor(`Training group`, levels = c("Training", "No training")),
Target = factor(Target, levels = c("Met", "Not Met"))
)
training_group_positions <- training_mosaic_data %>%
group_by(`Training group`) %>%
summarise(group_total = sum(count), .groups = "drop") %>%
mutate(
width = group_total / sum(group_total),
xmin = as.numeric(dplyr::lag(cumsum(width), default = 0)),
xmax = as.numeric(cumsum(width)),
xmid = (xmin + xmax) / 2
)
training_mosaic_data <- training_mosaic_data %>%
left_join(training_group_positions, by = "Training group") %>%
group_by(`Training group`) %>%
arrange(Target, .by_group = TRUE) %>%
mutate(
prop = count / group_total,
ymin = as.numeric(dplyr::lag(cumsum(prop), default = 0)),
ymax = as.numeric(cumsum(prop)),
ymid = (ymin + ymax) / 2
) %>%
ungroup()
training_mosaic_plot <- ggplot(training_mosaic_data) +
geom_rect(
aes(xmin = xmin, xmax = xmax, ymin = ymin, ymax = ymax, fill = Target),
colour = "white",
linewidth = 0.8
) +
geom_text(
aes(x = xmid, y = ymid, label = count),
family = "serif",
size = 4.1,
colour = "#111111"
) +
scale_x_continuous(
breaks = training_group_positions$xmid,
labels = training_group_positions$`Training group`,
expand = c(0, 0)
) +
scale_y_continuous(
labels = scales::percent_format(accuracy = 1),
expand = c(0, 0)
) +
scale_fill_manual(
values = c("Met" = "#6B9AC4", "Not Met" = "#C8C8C8"),
name = "Target status"
) +
labs(
x = "Training group",
y = "Within-group proportion",
title = "Training status and target attainment",
subtitle = "Block width reflects group size.\nBlock height shows the within-group proportion."
) +
theme_classic(base_size = 12) +
theme(
plot.title = element_text(size = 13),
plot.subtitle = element_text(size = 11),
legend.position = "top"
)
```
### Example: Fisher's Exact Test
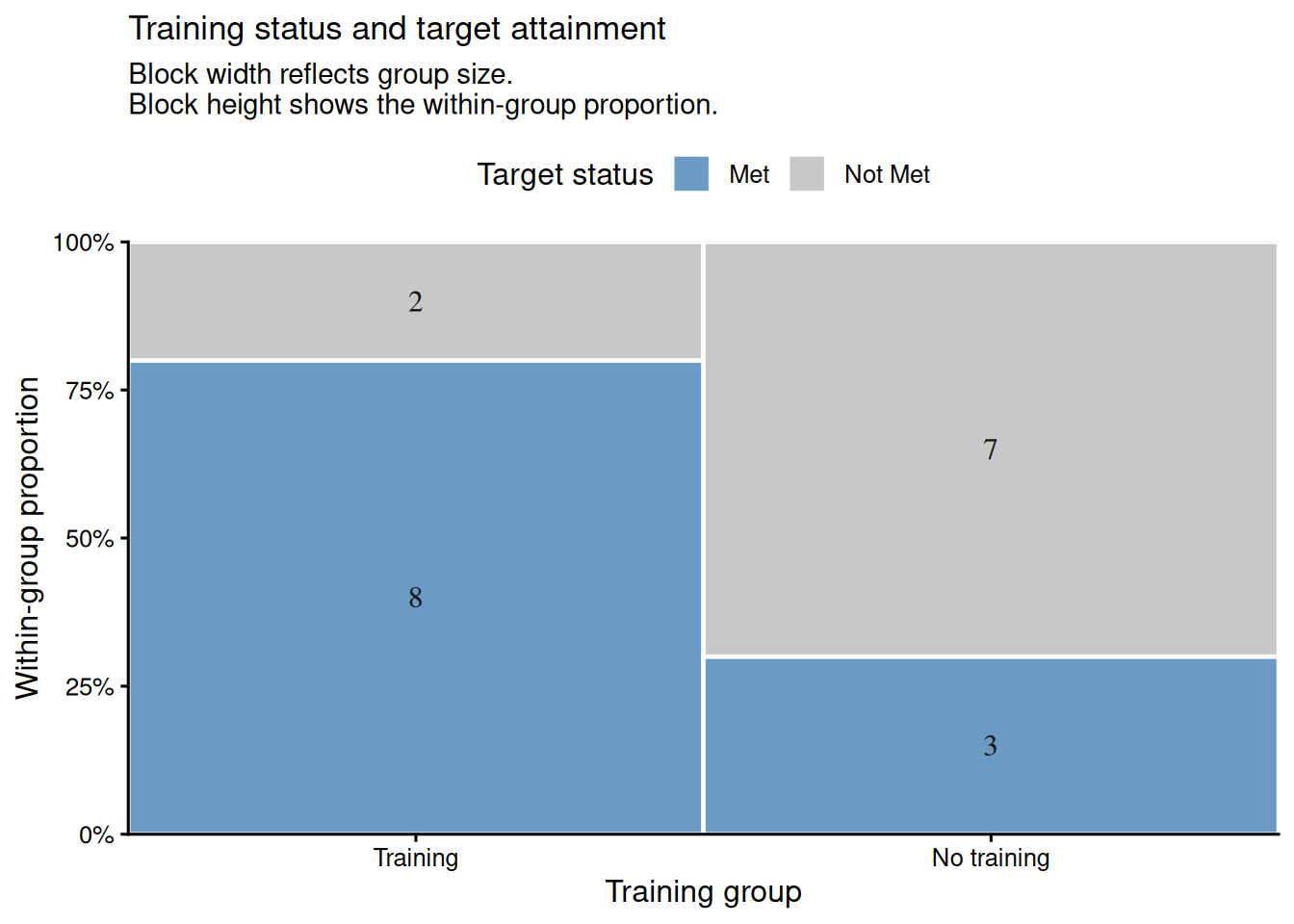
Suppose we are evaluating a new training intervention. Of 10 employees who received training, 8 met their performance target; of 10 who did not receive training, 3 met the target. We test whether training is associated with meeting the target.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-c-analysis-methods-chapter-10-exact-tests-and-resampling-methods-cell-1"}
#### Rendered Output
```{r}
#| label: chapter10-fisher-output-html
#| echo: false
#| results: asis
smallsamplelab_apa_table(
"10.1",
"Training outcomes by group",
training_display_table,
note = sprintf(
"The smallest expected cell count under independence is %.1f, so exact inference is preferable to the chi-square approximation.",
training_min_expected
)
)
```
#### Cell Code
```{webr-r}
#| context: interactive
training_table <- matrix(
c(8, 2, 3, 7),
nrow = 2,
byrow = TRUE,
dimnames = list(
Training = c("Yes", "No"),
Target = c("Met", "Not Met")
)
)
training_display_table <- tibble(
`Training group` = c("Training", "No training"),
`Met target` = c(8, 3),
`Did not meet target` = c(2, 7)
)
training_expected_counts <- suppressWarnings(chisq.test(training_table)$expected)
fisher_result <- fisher.test(training_table)
knitr::kable(training_display_table)
round(training_expected_counts, 1)
fisher_result
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: chapter10-fisher-output
#| results: asis
training_table <- matrix(
c(8, 2, 3, 7),
nrow = 2,
byrow = TRUE,
dimnames = list(
Training = c("Yes", "No"),
Target = c("Met", "Not Met")
)
)
training_display_table <- tibble(
`Training group` = c("Training", "No training"),
`Met target` = c(8, 3),
`Did not meet target` = c(2, 7)
)
training_expected_counts <- suppressWarnings(chisq.test(training_table)$expected)
fisher_result <- fisher.test(training_table)
smallsamplelab_apa_table(
"10.1",
"Training outcomes by group",
training_display_table,
note = sprintf(
"The smallest expected cell count under independence is %.1f, so exact inference is preferable to the chi-square approximation.",
min(training_expected_counts)
),
align = c("l", "r", "r")
)
```
::::
Fisher's exact test gives an odds ratio of `r sprintf("%.2f", unname(fisher_result$estimate))`, with a 95% confidence interval from `r sprintf("%.2f", fisher_result$conf.int[1])` to `r sprintf("%.2f", fisher_result$conf.int[2])`, and an exact two-sided p-value of `r chapter10_format_p(fisher_result$p.value)`. That odds ratio, despite the wide interval, points toward a meaningful association that a larger study would be well placed to assess.
#### Visualising the 2×2 Table
A mosaic-style plot helps readers see the same 2×2 structure graphically. The two training groups have equal width because each contains 10 employees, and the vertical split within each block shows the proportion who met the target. In this example, the training group has a visibly larger share of employees meeting the target.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-c-analysis-methods-chapter-10-exact-tests-and-resampling-methods-cell-1b"}
#### Rendered Output
```{r}
#| label: chapter10-fisher-mosaic-html
#| echo: false
#| fig-cap: "Figure 10.1: Mosaic-style plot of training status and target attainment. Block width reflects group size; block height within each group reflects the proportion meeting the target."
#| alt: "Mosaic-style plot of training status and target attainment. Block width reflects group size; block height within each group reflects the proportion meeting the target."
training_mosaic_plot
```
#### Cell Code
```{webr-r}
#| context: interactive
training_mosaic_data <- tibble(
`Training group` = c("Training", "Training", "No training", "No training"),
Target = c("Met", "Not Met", "Met", "Not Met"),
count = c(8, 2, 3, 7)
) %>%
mutate(
`Training group` = factor(`Training group`, levels = c("Training", "No training")),
Target = factor(Target, levels = c("Met", "Not Met"))
)
training_group_positions <- training_mosaic_data %>%
group_by(`Training group`) %>%
summarise(group_total = sum(count), .groups = "drop") %>%
mutate(
width = group_total / sum(group_total),
xmin = as.numeric(dplyr::lag(cumsum(width), default = 0)),
xmax = as.numeric(cumsum(width)),
xmid = (xmin + xmax) / 2
)
training_mosaic_data <- training_mosaic_data %>%
left_join(training_group_positions, by = "Training group") %>%
group_by(`Training group`) %>%
arrange(Target, .by_group = TRUE) %>%
mutate(
prop = count / group_total,
ymin = as.numeric(dplyr::lag(cumsum(prop), default = 0)),
ymax = as.numeric(cumsum(prop)),
ymid = (ymin + ymax) / 2
) %>%
ungroup()
ggplot(training_mosaic_data) +
geom_rect(
aes(xmin = xmin, xmax = xmax, ymin = ymin, ymax = ymax, fill = Target),
colour = "white",
linewidth = 0.8
) +
geom_text(
aes(x = xmid, y = ymid, label = count),
family = "serif",
size = 4.1,
colour = "#111111"
) +
scale_x_continuous(
breaks = training_group_positions$xmid,
labels = training_group_positions$`Training group`,
expand = c(0, 0)
) +
scale_y_continuous(
labels = scales::percent_format(accuracy = 1),
expand = c(0, 0)
) +
scale_fill_manual(
values = c("Met" = "#6B9AC4", "Not Met" = "#C8C8C8"),
name = "Target status"
) +
labs(
x = "Training group",
y = "Within-group proportion",
title = "Training status and target attainment",
subtitle = "Block width reflects group size.\nBlock height shows the within-group proportion."
) +
theme_classic(base_size = 12) +
theme(
plot.title = element_text(size = 13),
plot.subtitle = element_text(size = 11),
legend.position = "top"
)
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: chapter10-fisher-mosaic
#| fig-cap: "Figure 10.1: Mosaic-style plot of training status and target attainment. Block width reflects group size; block height within each group reflects the proportion meeting the target."
#| alt: "Mosaic-style plot of training status and target attainment. Block width reflects group size; block height within each group reflects the proportion meeting the target."
training_mosaic_plot
```
::::
### When Fisher's Exact Test Is Conservative
Fisher's exact test conditions on both row and column margins of the 2x2 table. In prospective studies, only the group sizes are fixed by design; conditioning on the observed outcome margin as well can make the test conservative. In case-control studies, both margins may align more closely with the sampling plan, though Fisher's conditioning remains stricter than the design itself.
- **Prospective study or trial**: group sizes are fixed by design, so one margin matches the design directly. Fisher's conditioning still also fixes the observed outcome margin, which is random and can make the test conservative.
- **Case-control study**: case and control totals are fixed by design, so the conditioning aligns more closely with the sampling plan, though Fisher's exact conditioning is still stricter than the design itself.
- **Cross-sectional or convenience sample**: neither margin is fixed by design, so the extra conditioning is most visible and conservatism is often easiest to see.
The practical implication is straightforward: a Fisher p-value can stay above 0.05 even when a less conservative exact method gives stronger evidence. Fisher's test remains valid, but the conditioning scheme affects how much power is available.
### Unconditional Exact Tests
The unconditional approach, developed by Barnard, does not require conditioning on both margins. Modern implementations therefore offer **unconditional exact tests** that condition only on the group sizes. These tests are often more powerful than Fisher's exact test when the margins are not fixed by design.
In R, the `exact2x2` package provides unconditional exact procedures through `uncondExact2x2()` [@fay2010]. Install it with `install.packages("exact2x2")` if it is not already available. The score version shown below is Barnard-style in spirit and compares the observed proportions while conditioning only on the group sizes.
```{r}
#| include: false
library(exact2x2)
midp_result <- exact2x2(training_table, alternative = "two.sided", midp = TRUE)
barnard_result <- uncondExact2x2(
x1 = 8,
n1 = 10,
x2 = 3,
n2 = 10,
parmtype = "difference",
alternative = "two.sided",
method = "score",
conf.int = TRUE
)
# uncondExact2x2() returns p2 - p1, so reverse the sign to report training minus no training.
barnard_risk_difference <- unname(-barnard_result$estimate)
barnard_risk_difference_ci <- c(-barnard_result$conf.int[2], -barnard_result$conf.int[1])
barnard_note <- sprintf(
"The unconditional exact score test estimates a risk difference of %.2f, with a 95%% confidence interval from %.2f to %.2f.",
barnard_risk_difference,
barnard_risk_difference_ci[1],
barnard_risk_difference_ci[2]
)
exact_comparison_table <- tibble(
Method = c(
"Fisher's exact test",
"Fisher's exact test with mid-p correction",
"Unconditional exact score test"
),
Conditioning = c(
"Conditions on both margins",
"Conditions on both margins, then applies a mid-p adjustment",
"Conditions on group sizes only"
),
`Two-sided p-value` = c(
chapter10_format_p(fisher_result$p.value),
chapter10_format_p(midp_result$p.value),
chapter10_format_p(barnard_result$p.value)
)
)
```
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-c-analysis-methods-chapter-10-exact-tests-and-resampling-methods-cell-2"}
#### Rendered Output
```{r}
#| label: chapter10-barnard-output-html
#| echo: false
#| results: asis
smallsamplelab_apa_table(
"10.2",
"Comparison of exact p-values for the training example",
exact_comparison_table,
note = barnard_note
)
```
#### Cell Code
```{webr-r}
#| context: interactive
library(exact2x2)
midp_result <- exact2x2(training_table, alternative = "two.sided", midp = TRUE)
barnard_result <- uncondExact2x2(
x1 = 8,
n1 = 10,
x2 = 3,
n2 = 10,
parmtype = "difference",
alternative = "two.sided",
method = "score",
conf.int = TRUE
)
# uncondExact2x2() returns p2 - p1, so reverse the sign to report training minus no training.
barnard_risk_difference <- unname(-barnard_result$estimate)
barnard_risk_difference_ci <- c(-barnard_result$conf.int[2], -barnard_result$conf.int[1])
exact_comparison_table <- tibble(
Method = c(
"Fisher's exact test",
"Fisher's exact test with mid-p correction",
"Unconditional exact score test"
),
Conditioning = c(
"Conditions on both margins",
"Conditions on both margins, then applies a mid-p adjustment",
"Conditions on group sizes only"
),
`Two-sided p-value` = c(
chapter10_format_p(fisher_result$p.value),
chapter10_format_p(midp_result$p.value),
chapter10_format_p(barnard_result$p.value)
)
)
knitr::kable(exact_comparison_table)
barnard_result
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: chapter10-barnard-output
#| results: asis
library(exact2x2)
midp_result <- exact2x2(training_table, alternative = "two.sided", midp = TRUE)
barnard_result <- uncondExact2x2(
x1 = 8,
n1 = 10,
x2 = 3,
n2 = 10,
parmtype = "difference",
alternative = "two.sided",
method = "score",
conf.int = TRUE
)
# uncondExact2x2() returns p2 - p1, so reverse the sign to report training minus no training.
barnard_risk_difference <- unname(-barnard_result$estimate)
barnard_risk_difference_ci <- c(-barnard_result$conf.int[2], -barnard_result$conf.int[1])
exact_comparison_table <- tibble(
Method = c(
"Fisher's exact test",
"Fisher's exact test with mid-p correction",
"Unconditional exact score test"
),
Conditioning = c(
"Conditions on both margins",
"Conditions on both margins, then applies a mid-p adjustment",
"Conditions on group sizes only"
),
`Two-sided p-value` = c(
chapter10_format_p(fisher_result$p.value),
chapter10_format_p(midp_result$p.value),
chapter10_format_p(barnard_result$p.value)
)
)
smallsamplelab_apa_table(
"10.2",
"Comparison of exact p-values for the training example",
exact_comparison_table,
note = barnard_note,
align = c("l", "l", "r")
)
```
::::
For this table, Fisher's exact p-value is `r chapter10_format_p(fisher_result$p.value)`, whereas the mid-p correction gives `r chapter10_format_p(midp_result$p.value)` and the unconditional exact score test gives `r chapter10_format_p(barnard_result$p.value)`. The unconditional method is less conservative here because it does not condition on both margins. Reported as training minus no training, the estimated risk difference is `r sprintf("%.2f", barnard_risk_difference)`, with a 95% confidence interval from `r sprintf("%.2f", barnard_risk_difference_ci[1])` to `r sprintf("%.2f", barnard_risk_difference_ci[2])`.
### Mid-p Corrections
Mid-p corrections reduce conservatism by subtracting half the probability of the observed table from the tail area. This yields a test that does not guarantee Type I error control at the nominal level in every configuration. Report mid-p results as a sensitivity analysis alongside the standard Fisher test rather than as a replacement default.
::: {.callout-warning}
## Common Misconception: "Exact" Means "Automatically Best"
**Myth**: "If a test is exact, it must be the most appropriate or most informative option."
**Reality**: Exactness describes how the p-value is computed under the null hypothesis. The appropriate conditioning scheme still depends on the design. In the training example above, Fisher's exact test gives `r chapter10_format_p(fisher_result$p.value)`, while the mid-p and unconditional exact alternatives give `r chapter10_format_p(midp_result$p.value)` and `r chapter10_format_p(barnard_result$p.value)` respectively.
**Lesson**:
1. Exact computation and design appropriateness are separate questions.
2. Fisher's exact test is a strong default for confirmatory sparse 2×2 analyses.
3. Mid-p and unconditional exact tests are useful sensitivity checks when Fisher's conditioning may be overly restrictive.
4. With very small samples, the effect estimate and the underlying table can be more informative than a p-value alone.
:::
### Exact Binomial Test
The exact binomial test assesses whether the observed number of successes is compatible with a hypothesised success probability. It is appropriate for a single binary outcome when the null benchmark is known in advance or supplied by design.
```{r}
#| include: false
binom_result <- binom.test(x = 13, n = 15, p = 0.70, alternative = "two.sided")
binom_summary_table <- tibble(
`Observed successes` = "13 of 15",
`Observed proportion` = sprintf("%.3f", 13 / 15),
`Hypothesised proportion` = sprintf("%.2f", 0.70),
`95% exact CI` = sprintf("%.2f to %.2f", binom_result$conf.int[1], binom_result$conf.int[2]),
`Exact p-value` = chapter10_format_p(binom_result$p.value)
)
```
### Example: Exact Binomial Test
A clinic claims that 70% of patients improve with standard care. In a small audit of 15 patients, 13 improved. We test whether the observed proportion is consistent with the clinic's claim.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-c-analysis-methods-chapter-10-exact-tests-and-resampling-methods-cell-3"}
#### Rendered Output
```{r}
#| label: chapter10-binom-output-html
#| echo: false
#| results: asis
smallsamplelab_apa_table(
"10.3",
"Exact binomial test summary",
binom_summary_table
)
```
#### Cell Code
```{webr-r}
#| context: interactive
binom_result <- binom.test(x = 13, n = 15, p = 0.70, alternative = "two.sided")
binom_summary_table <- tibble(
`Observed successes` = "13 of 15",
`Observed proportion` = sprintf("%.3f", 13 / 15),
`Hypothesised proportion` = sprintf("%.2f", 0.70),
`95% exact CI` = sprintf("%.2f to %.2f", binom_result$conf.int[1], binom_result$conf.int[2]),
`Exact p-value` = chapter10_format_p(binom_result$p.value)
)
knitr::kable(binom_summary_table)
binom_result
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: chapter10-binom-output
#| results: asis
binom_result <- binom.test(x = 13, n = 15, p = 0.70, alternative = "two.sided")
binom_summary_table <- tibble(
`Observed successes` = "13 of 15",
`Observed proportion` = sprintf("%.3f", 13 / 15),
`Hypothesised proportion` = sprintf("%.2f", 0.70),
`95% exact CI` = sprintf("%.2f to %.2f", binom_result$conf.int[1], binom_result$conf.int[2]),
`Exact p-value` = chapter10_format_p(binom_result$p.value)
)
smallsamplelab_apa_table(
"10.3",
"Exact binomial test summary",
binom_summary_table,
align = c("l", "r", "r", "l", "r")
)
```
::::
The observed improvement proportion is `r sprintf("%.3f", 13 / 15)`, and the exact binomial p-value is `r chapter10_format_p(binom_result$p.value)`. With such a small audit, the data remain compatible with the clinic's claimed 70% rate. The 95% exact confidence interval from `r sprintf("%.2f", binom_result$conf.int[1])` to `r sprintf("%.2f", binom_result$conf.int[2])` is wide enough to include both the claimed value and substantially higher improvement rates.
### Exact Poisson Test
The exact Poisson test is used for count data when the quantity of interest is the number of events in a fixed amount of time, area, or exposure. It tests whether an observed count is consistent with a specified Poisson rate.
```{r}
#| include: false
poisson_observed_count <- 32
poisson_batches <- 8
poisson_expected_rate <- 3
poisson_result <- poisson.test(
x = poisson_observed_count,
T = poisson_batches,
r = poisson_expected_rate,
alternative = "two.sided"
)
poisson_summary_table <- tibble(
`Observed count` = sprintf("%d defects across %d batches", poisson_observed_count, poisson_batches),
`Observed rate` = sprintf("%.2f per batch", poisson_observed_count / poisson_batches),
`Hypothesised rate` = sprintf("%.2f per batch", poisson_expected_rate),
`95% exact CI` = sprintf("%.2f to %.2f", poisson_result$conf.int[1], poisson_result$conf.int[2]),
`Exact p-value` = chapter10_format_p(poisson_result$p.value)
)
```
### Example: Exact Poisson Test
A manufacturing process is expected to produce `r poisson_expected_rate` defects per batch on average. In a random sample of `r poisson_batches` batches, we observe `r poisson_observed_count` total defects. We test whether the observed rate is consistent with the expected rate of `r poisson_expected_rate` per batch.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-c-analysis-methods-chapter-10-exact-tests-and-resampling-methods-cell-4"}
#### Rendered Output
```{r}
#| label: chapter10-poisson-output-html
#| echo: false
#| results: asis
smallsamplelab_apa_table(
"10.4",
"Exact Poisson test summary",
poisson_summary_table
)
```
#### Cell Code
```{webr-r}
#| context: interactive
poisson_observed_count <- 32
poisson_batches <- 8
poisson_expected_rate <- 3
poisson_result <- poisson.test(
x = poisson_observed_count,
T = poisson_batches,
r = poisson_expected_rate,
alternative = "two.sided"
)
poisson_summary_table <- tibble(
`Observed count` = sprintf("%d defects across %d batches", poisson_observed_count, poisson_batches),
`Observed rate` = sprintf("%.2f per batch", poisson_observed_count / poisson_batches),
`Hypothesised rate` = sprintf("%.2f per batch", poisson_expected_rate),
`95% exact CI` = sprintf("%.2f to %.2f", poisson_result$conf.int[1], poisson_result$conf.int[2]),
`Exact p-value` = chapter10_format_p(poisson_result$p.value)
)
knitr::kable(poisson_summary_table)
poisson_result
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: chapter10-poisson-output
#| results: asis
poisson_observed_count <- 32
poisson_batches <- 8
poisson_expected_rate <- 3
poisson_result <- poisson.test(
x = poisson_observed_count,
T = poisson_batches,
r = poisson_expected_rate,
alternative = "two.sided"
)
poisson_summary_table <- tibble(
`Observed count` = sprintf("%d defects across %d batches", poisson_observed_count, poisson_batches),
`Observed rate` = sprintf("%.2f per batch", poisson_observed_count / poisson_batches),
`Hypothesised rate` = sprintf("%.2f per batch", poisson_expected_rate),
`95% exact CI` = sprintf("%.2f to %.2f", poisson_result$conf.int[1], poisson_result$conf.int[2]),
`Exact p-value` = chapter10_format_p(poisson_result$p.value)
)
smallsamplelab_apa_table(
"10.4",
"Exact Poisson test summary",
poisson_summary_table,
align = c("l", "r", "l", "l", "r")
)
```
::::
The observed defect rate is `r sprintf("%.2f", 32 / 8)` per batch, compared with the hypothesised rate of 3.00. The exact Poisson p-value is `r chapter10_format_p(poisson_result$p.value)`, so the data are broadly consistent with the expected defect rate. The 95% exact interval from `r sprintf("%.2f", poisson_result$conf.int[1])` to `r sprintf("%.2f", poisson_result$conf.int[2])` still includes 3.00.
### Permutation Tests
Permutation tests compare the observed statistic with the distribution obtained by rearranging group labels under the null hypothesis. When every possible reallocation is enumerated, the resulting p-value is exact under the null. When only a large random sample of reallocations is used, the result is a Monte Carlo approximation. For two groups of size n1 and n2, the number of reallocations is (n1 + n2)! / (n1! n2!), equivalent to choosing n1 observations from the pooled sample. If this number is computationally feasible, for example below 100,000, enumerate all permutations; otherwise use a large Monte Carlo sample such as B = 10,000 and report both B and the random seed.
The key assumption is **exchangeability under the null hypothesis**: if the null is true, the joint distribution of the data remains the same under any reassignment of labels. That makes permutation tests especially attractive for small-sample comparisons where a parametric model would be difficult to justify.
```{r}
#| include: false
method_a <- c(78, 82, 75, 88, 71, 85, 80, 83)
method_b <- c(68, 72, 76, 75, 79, 69, 73, 74)
all_scores <- c(method_a, method_b)
obs_diff <- mean(method_a) - mean(method_b)
perm_combos <- combn(seq_along(all_scores), length(method_a))
perm_diffs <- apply(
perm_combos,
2,
function(idx) mean(all_scores[idx]) - mean(all_scores[-idx])
)
perm_p_value <- mean(abs(perm_diffs) >= abs(obs_diff))
perm_plot_data <- tibble(diff = perm_diffs)
perm_plot <- ggplot(perm_plot_data, aes(x = diff)) +
geom_histogram(
binwidth = 1,
boundary = 0,
colour = "white",
fill = "#A7D3F2"
) +
geom_vline(xintercept = obs_diff, colour = "#C0392B", linewidth = 1) +
geom_vline(xintercept = -obs_diff, colour = "#C0392B", linewidth = 1, linetype = "dashed") +
labs(
x = "Difference in means (A - B)",
y = "Number of reallocations",
title = "Exact permutation distribution of the mean difference",
subtitle = "Red lines mark the observed difference\nand its mirror image under a two-sided test."
) +
theme_classic(base_size = 12) +
theme(
plot.title = element_text(size = 13),
plot.subtitle = element_text(size = 11)
)
```
### Example: Permutation Test for Difference in Means
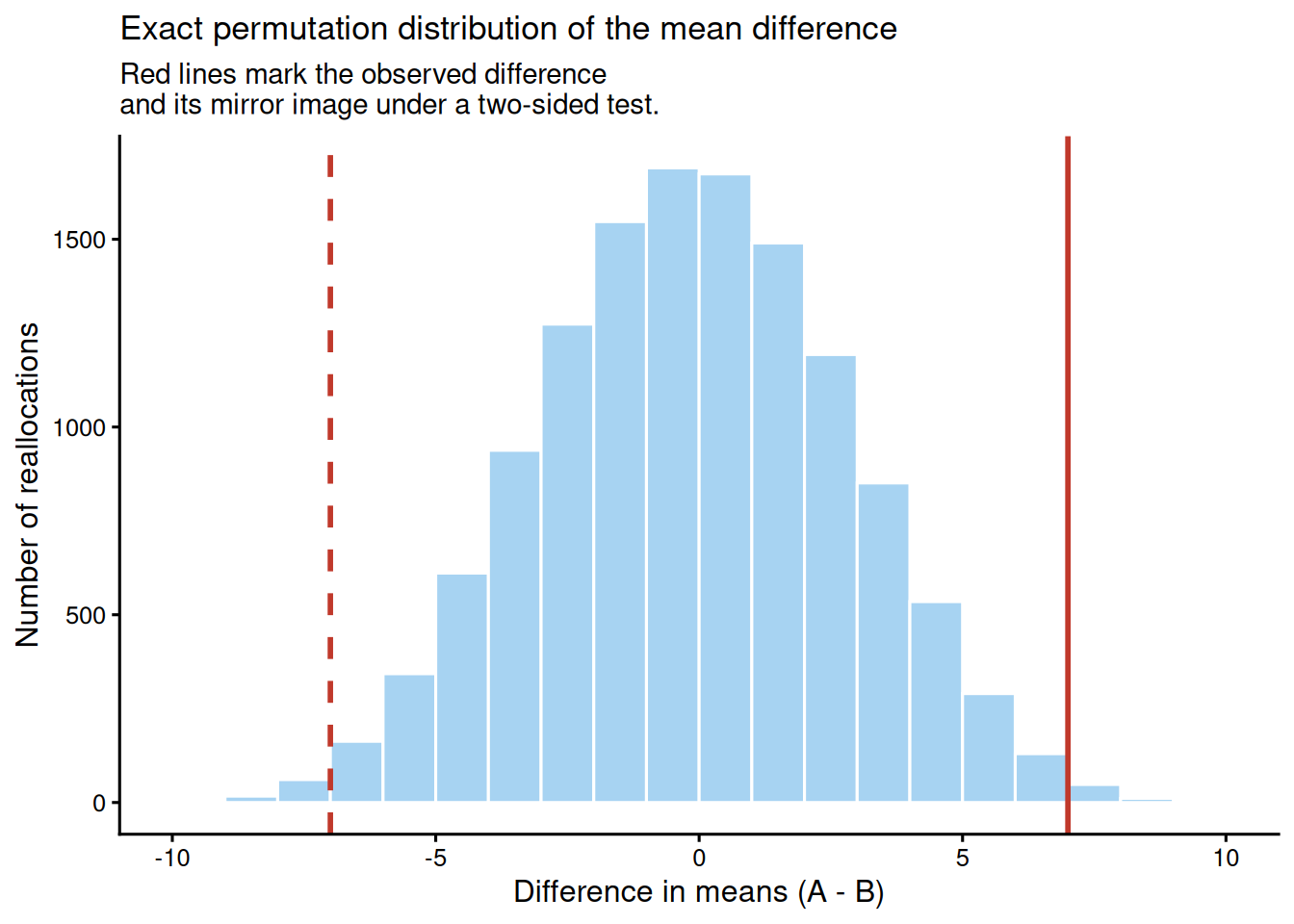
We compare test scores between two teaching methods with small groups of 8 students each. The outcome is continuous, but with such a small sample it is reasonable to avoid normality assumptions and to test the mean difference by permutation.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-c-analysis-methods-chapter-10-exact-tests-and-resampling-methods-cell-5"}
#### Rendered Output
```{r}
#| label: chapter10-permutation-output-html
#| echo: false
#| fig-cap: "Figure 10.2: Exact permutation distribution for the mean difference between teaching methods."
#| alt: "Exact permutation distribution for the mean difference between teaching methods."
perm_plot
```
#### Cell Code
```{webr-r}
#| context: interactive
method_a <- c(78, 82, 75, 88, 71, 85, 80, 83)
method_b <- c(68, 72, 76, 75, 79, 69, 73, 74)
all_scores <- c(method_a, method_b)
obs_diff <- mean(method_a) - mean(method_b)
perm_combos <- combn(seq_along(all_scores), length(method_a))
perm_diffs <- apply(
perm_combos,
2,
function(idx) mean(all_scores[idx]) - mean(all_scores[-idx])
)
perm_p_value <- mean(abs(perm_diffs) >= abs(obs_diff))
perm_plot_data <- tibble(diff = perm_diffs)
ggplot(perm_plot_data, aes(x = diff)) +
geom_histogram(
binwidth = 1,
boundary = 0,
colour = "white",
fill = "#A7D3F2"
) +
geom_vline(xintercept = obs_diff, colour = "#C0392B", linewidth = 1) +
geom_vline(xintercept = -obs_diff, colour = "#C0392B", linewidth = 1, linetype = "dashed") +
labs(
x = "Difference in means (A - B)",
y = "Number of reallocations",
title = "Exact permutation distribution of the mean difference",
subtitle = "Red lines mark the observed difference\nand its mirror image under a two-sided test."
) +
theme_classic(base_size = 12)
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: chapter10-permutation-output
#| fig-cap: "Figure 10.2: Exact permutation distribution for the mean difference between teaching methods."
#| alt: "Exact permutation distribution for the mean difference between teaching methods."
method_a <- c(78, 82, 75, 88, 71, 85, 80, 83)
method_b <- c(68, 72, 76, 75, 79, 69, 73, 74)
all_scores <- c(method_a, method_b)
obs_diff <- mean(method_a) - mean(method_b)
perm_combos <- combn(seq_along(all_scores), length(method_a))
perm_diffs <- apply(
perm_combos,
2,
function(idx) mean(all_scores[idx]) - mean(all_scores[-idx])
)
perm_p_value <- mean(abs(perm_diffs) >= abs(obs_diff))
perm_plot_data <- tibble(diff = perm_diffs)
ggplot(perm_plot_data, aes(x = diff)) +
geom_histogram(
binwidth = 1,
boundary = 0,
colour = "white",
fill = "#A7D3F2"
) +
geom_vline(xintercept = obs_diff, colour = "#C0392B", linewidth = 1) +
geom_vline(xintercept = -obs_diff, colour = "#C0392B", linewidth = 1, linetype = "dashed") +
labs(
x = "Difference in means (A - B)",
y = "Number of reallocations",
title = "Exact permutation distribution of the mean difference",
subtitle = "Red lines mark the observed difference\nand its mirror image under a two-sided test."
) +
theme_classic(base_size = 12)
```
::::
Across all `r format(length(perm_diffs), big.mark = ",")` possible reallocations, the observed mean difference is `r sprintf("%.2f", obs_diff)` points and the exact two-sided permutation p-value is `r chapter10_format_p(perm_p_value, digits = 4)`. Four-decimal precision is shown here because the exact enumeration produces a very small non-zero p-value. Because every possible label assignment is included, this result is exact rather than Monte Carlo. The figure makes the logic visible: the observed difference sits at the extreme tail of the permutation distribution.
### Bootstrap Confidence Intervals
Bootstrap resampling constructs confidence intervals by repeatedly drawing samples with replacement from the observed data and recalculating the statistic of interest. The resulting bootstrap distribution approximates the sampling distribution of that statistic.
This chapter uses the **nonparametric bootstrap**, which resamples directly from the observed data. A **parametric bootstrap** simulates new datasets from a fitted parametric model, so its validity depends entirely on how well that model represents the data. For small-sample work, the nonparametric bootstrap is often the safer default when a model-free interval is desired.
```{r}
#| include: false
library(boot)
recovery_times <- c(12, 15, 14, 18, 16, 13, 17, 19, 14, 15, 20, 16, 15, 18, 17)
sample_median <- median(recovery_times)
median_fun <- function(data, indices) {
median(data[indices])
}
set.seed(2025)
boot_result <- boot(data = recovery_times, statistic = median_fun, R = 2000)
boot_ci <- boot.ci(boot_result, conf = 0.95, type = c("perc", "bca"))
boot_percentile_ci <- boot_ci$percent[4:5]
boot_bca_ci <- boot_ci$bca[4:5]
bootstrap_summary_table <- tibble(
Statistic = "Median recovery time",
Estimate = sprintf("%.0f days", sample_median),
`95% percentile bootstrap CI` = sprintf(
"%.0f to %.0f days",
boot_percentile_ci[1],
boot_percentile_ci[2]
),
`95% BCa bootstrap CI` = sprintf(
"%.0f to %.0f days",
boot_bca_ci[1],
boot_bca_ci[2]
),
Resamples = format(boot_result$R, big.mark = ",")
)
```
### Example: Bootstrap CI for the Median
We estimate the median recovery time for a small sample of patients and construct a 95% bootstrap confidence interval.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-c-analysis-methods-chapter-10-exact-tests-and-resampling-methods-cell-6"}
#### Rendered Output
```{r}
#| label: chapter10-bootstrap-output-html
#| echo: false
#| results: asis
smallsamplelab_apa_table(
"10.5",
"Bootstrap summary for median recovery time",
bootstrap_summary_table
)
```
#### Cell Code
```{webr-r}
#| context: interactive
library(boot)
recovery_times <- c(12, 15, 14, 18, 16, 13, 17, 19, 14, 15, 20, 16, 15, 18, 17)
sample_median <- median(recovery_times)
median_fun <- function(data, indices) {
median(data[indices])
}
set.seed(2025)
boot_result <- boot(data = recovery_times, statistic = median_fun, R = 2000)
boot_ci <- boot.ci(boot_result, conf = 0.95, type = c("perc", "bca"))
bootstrap_summary_table <- tibble(
Statistic = "Median recovery time",
Estimate = sprintf("%.0f days", sample_median),
`95% percentile bootstrap CI` = sprintf(
"%.0f to %.0f days",
boot_ci$percent[4],
boot_ci$percent[5]
),
`95% BCa bootstrap CI` = sprintf(
"%.0f to %.0f days",
boot_ci$bca[4],
boot_ci$bca[5]
),
Resamples = format(boot_result$R, big.mark = ",")
)
knitr::kable(bootstrap_summary_table)
boot_ci
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: chapter10-bootstrap-output
#| results: asis
library(boot)
recovery_times <- c(12, 15, 14, 18, 16, 13, 17, 19, 14, 15, 20, 16, 15, 18, 17)
sample_median <- median(recovery_times)
median_fun <- function(data, indices) {
median(data[indices])
}
set.seed(2025)
boot_result <- boot(data = recovery_times, statistic = median_fun, R = 2000)
boot_ci <- boot.ci(boot_result, conf = 0.95, type = c("perc", "bca"))
bootstrap_summary_table <- tibble(
Statistic = "Median recovery time",
Estimate = sprintf("%.0f days", sample_median),
`95% percentile bootstrap CI` = sprintf(
"%.0f to %.0f days",
boot_ci$percent[4],
boot_ci$percent[5]
),
`95% BCa bootstrap CI` = sprintf(
"%.0f to %.0f days",
boot_ci$bca[4],
boot_ci$bca[5]
),
Resamples = format(boot_result$R, big.mark = ",")
)
smallsamplelab_apa_table(
"10.5",
"Bootstrap summary for median recovery time",
bootstrap_summary_table,
align = c("l", "r", "l", "l", "r")
)
```
::::
The sample median recovery time is `r sprintf("%.0f", sample_median)` days. The 95% percentile bootstrap interval runs from `r sprintf("%.0f", boot_percentile_ci[1])` to `r sprintf("%.0f", boot_percentile_ci[2])` days, and the BCa interval runs from `r sprintf("%.0f", boot_bca_ci[1])` to `r sprintf("%.0f", boot_bca_ci[2])` days. Both intervals summarise uncertainty around the median without requiring a normal-theory standard error formula. Percentile intervals are simple; BCa intervals adjust for bias and acceleration in the bootstrap distribution [@efron1993]. With n < 30, BCa intervals can be unstable or fail when the statistic has many ties, so report the interval type and inspect the bootstrap distribution rather than treating BCa as automatically superior.
### Key Takeaways
Exact tests are most valuable when the data are sparse enough that large-sample approximations become questionable. Fisher's exact test is a strong default for sparse 2×2 tables, but mid-p and unconditional exact procedures can serve as useful sensitivity checks when Fisher's conditioning is stronger than the design requires. Exact binomial and exact Poisson tests extend the same logic to single proportions and sparse event rates, while permutation tests rely on exchangeability and can be exact when every possible reallocation is enumerated. Bootstrap intervals are especially useful for statistics such as medians that lack simple analytic standard errors, provided the resampling procedure is reported transparently.
### Self-Assessment Quiz
Test your understanding of exact tests and resampling methods from Chapter 10.
```{r}
#| echo: false
#| results: asis
quiz_helpers_path <- normalizePath(file.path(dirname(knitr::current_input(dir = TRUE)), "..", "R", "quiz_helpers.R"), mustWork = FALSE)
if (file.exists(quiz_helpers_path)) {
source(quiz_helpers_path)
} else {
stop("Quiz helper file not found at: ", quiz_helpers_path, ". Please ensure 'quiz_helpers.R' exists in the R directory.")
}
smallsamplelab_render_quiz(list(
list(
prompt = "When is Fisher's exact test especially appropriate?",
options = c("When every expected cell count is comfortably above 20", "When a 2×2 table has sparse counts and exact inference is preferred", "When the outcome is continuous and normally distributed", "When the goal is estimating a time-series trend"),
answer = 2L,
explanation = "Fisher's exact test is designed for sparse 2×2 tables where exact inference is preferred to the chi-square approximation. The chapter introduces it as the standard conditional test when some expected cell counts are small."
),
list(
prompt = "Why can Fisher's exact test be conservative in some settings?",
options = c("It ignores the observed table", "It conditions on both margins, which can make rejection harder than necessary when both are not fixed by design", "It always assumes normality", "It uses too many bootstrap resamples"),
answer = 2L,
explanation = "The chapter explains that Fisher's test conditions on both margins. When the design leaves one margin free, that conditioning can make the test more conservative and reduce power."
),
list(
prompt = "When might Barnard-style unconditional tests or a mid-p correction be considered?",
options = c("When you need a less conservative sensitivity check for sparse 2×2 data", "When the outcome is continuous and homoscedastic", "When all expected counts exceed 100", "When fitting Bayesian regression with weakly informative priors"),
answer = 1L, # option A is correct here
explanation = "The chapter presents mid-p and unconditional exact procedures as less conservative alternatives or sensitivity checks for sparse 2×2 tables, especially when Fisher's conditioning may be overly restrictive."
),
list(
prompt = "What does the exact binomial test evaluate?",
options = c("Whether two means differ after random permutation", "Whether an observed number of successes is consistent with a hypothesised proportion", "Whether a count process is overdispersed relative to Poisson", "Whether a regression model has converged"),
answer = 2L,
explanation = "The exact binomial test is for a single-sample binary outcome. It compares the observed number of successes with a fixed benchmark proportion without relying on large-sample approximations."
),
list(
prompt = "What is the exact Poisson test used for in this chapter?",
options = c("Comparing two continuous variables with tied ranks", "Testing whether an observed event count or rate is consistent with a specified Poisson rate", "Selecting variables in penalised regression", "Checking MCMC convergence"),
answer = 2L,
explanation = "The chapter uses the exact Poisson test for sparse count data when the interest is whether an observed count or event rate differs from a known benchmark."
),
list(
prompt = "What key assumption makes a permutation test valid under the null hypothesis?",
options = c("Exchangeability of observations or labels under the null", "Normal residuals with equal variance", "At least 100 observations per group", "A Poisson-distributed outcome"),
answer = 1L, # option A is correct here
explanation = "Permutation tests rely on exchangeability under the null hypothesis. If the joint distribution of the data remains the same under any reassignment of labels, the permutation reference distribution is valid."
),
list(
prompt = "Why is the bootstrap especially useful in small-sample work?",
options = c("It guarantees narrow confidence intervals", "It can approximate uncertainty for statistics like medians when closed-form standard errors are unavailable", "It removes the need to report uncertainty", "It is only valid for normal data"),
answer = 2L,
explanation = "The bootstrap is useful when the statistic of interest does not have a simple analytic standard error or when a model-free interval is preferred. The chapter's median example illustrates this directly."
),
list(
prompt = "Which reporting detail is most important to include for resampling procedures?",
options = c("Only whether the result was significant", "The number of resamples or permutations, the statistic used, and enough detail to reproduce the procedure", "Only the sample size", "Only the largest resampled statistic"),
answer = 2L,
explanation = "The chapter states that resampling analyses should report the statistic being resampled, the number of permutations or bootstrap resamples, the random seed, and whether the result is exact or Monte Carlo."
)
))
```