```{r}
#| include: false
suppressPackageStartupMessages(library(tidyverse))
suppressPackageStartupMessages(library(knitr))
suppressPackageStartupMessages(library(htmltools))
source(normalizePath(file.path(dirname(knitr::current_input(dir = TRUE)), "..", "R", "chapter_helpers.R"), mustWork = TRUE))
```
:::: {.content-visible when-format="html"}
```{webr-r}
#| context: setup
#| include: false
#| echo: false
smallsamplelab_apa_table <- function(number, caption, data, note = NULL, align = NULL, ...) {
print(data)
if (!is.null(note)) message(note)
invisible(data)
}
smalln_mcar_table <- function(data) {
missing_cells <- sum(is.na(data))
data.frame(
Test = "Little's MCAR test (illustrative browser fallback)",
`Missing cells` = missing_cells,
`Missing percent` = round(100 * missing_cells / length(data), 1),
check.names = FALSE
)
}
smalln_missing_var_plot <- function(data) {
missing_percent <- colMeans(is.na(data)) * 100
barplot(missing_percent, ylab = "Missing (%)", las = 2, col = "#2E7D6E")
}
smalln_missing_matrix_plot <- function(data) {
missing_matrix <- t(is.na(as.data.frame(data)))
image(missing_matrix[nrow(missing_matrix):1, ], axes = FALSE, col = c("white", "#2E7D6E"))
axis(1, at = seq(0, 1, length.out = nrow(data)), labels = seq_len(nrow(data)), cex.axis = 0.7)
axis(2, at = seq(0, 1, length.out = ncol(data)), labels = rev(names(data)), las = 2, cex.axis = 0.7)
}
```
::::
# Chapter 8: Handling Missing Data in Small Samples
### Learning Objectives
By the end of this chapter, you will be able to distinguish MCAR, MAR, and MNAR mechanisms, describe missingness patterns clearly, judge when complete-case analysis or multiple imputation is defensible in a small sample, and report missing-data decisions with the transparency needed for readers to assess their consequences.
### The Challenge of Missing Data in Small Samples
Missing data are common in applied research. Participants skip survey questions, drop out of longitudinal studies, or provide incomplete records. With large samples, modern methods (multiple imputation, full information maximum likelihood) can handle substantial missingness without excessive bias. With small samples, however, missing data pose severe problems. Even a few missing observations can substantially reduce effective sample size and statistical power.
Missing data methods rely on large-sample asymptotics and may be unstable or inappropriate when samples are very small (n < 30) or missingness is extensive (> 20%). In such cases, prevention (minimise missingness through careful design) and transparency (report missingness patterns and sensitivity analyses) are more important than sophisticated imputation.
### Types of Missingness
The standard missingness framework distinguishes between three mechanisms [@rubin1987; @vanbuuren2018]. **MCAR (Missing Completely At Random)** means missingness is unrelated to any observed or unobserved variable, as when a survey page disappears because of a random software glitch. This condition is conceptually simple but rare in practice. **MAR (Missing At Random)** means missingness depends on observed variables but not on the missing values themselves once those observed variables are taken into account. For example, older participants may be more likely to skip a technology question, but conditional on age the missingness is otherwise random. **MNAR (Missing Not At Random)** is the hardest case because missingness depends on the unobserved values themselves, as when participants with severe depression are especially likely to drop out. That possibility cannot usually be resolved from the observed data alone and therefore requires sensitivity analysis or explicit modelling assumptions.
### Describing Missingness Patterns
Before choosing a handling strategy, describe the pattern of missingness in plain terms. Report how many observations are missing on each variable, whether the missing values are concentrated in particular participants or particular variables, and whether incomplete cases differ from complete cases on observed characteristics that might make MAR more plausible than MCAR.
### Example Dataset for Diagnostics
To demonstrate the handling strategies in this chapter, we simulate a small dataset with missing values on `satisfaction` and `performance`. Table 8.1 shows the first ten rows so the missing-data pattern is visible before the formal diagnostics begin.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-b-data-collection-chapter-12-handling-missing-data-in-small-samples-cell-1"}
#### Rendered Output
```{r}
#| label: part-b-chunk-14-html
#| echo: false
library(tidyverse)
set.seed(2025)
# Realistic age distribution and MAR missingness:
# performance is more likely to be missing for older participants
age_raw <- round(rnorm(25, mean = 38, sd = 12))
age_vals <- pmax(18L, pmin(65L, age_raw))
perf_full <- round(40 + 0.6 * age_vals + rnorm(25, 0, 10))
miss_prob <- plogis((age_vals - 42) / 10)
perf_obs <- ifelse(runif(25) < miss_prob * 0.55, NA_real_, perf_full)
study_data <- tibble(
participant = 1:25,
age = age_vals,
satisfaction = sample(c(3:7, NA), 25, replace = TRUE,
prob = c(0.15, 0.2, 0.25, 0.2, 0.15, 0.05)),
performance = perf_obs
)
study_data_display <- study_data %>%
mutate(across(c(satisfaction, performance), ~ if_else(is.na(.x), "NA", as.character(.x)))) %>%
slice(1:10)
smallsamplelab_apa_table(
"8.1",
"Simulated study dataset with missing values (rows 1 to 10)",
study_data_display,
note = "The full simulated dataset contains 25 participants; the excerpt is shown to illustrate the missing values before diagnosis.",
align = c("r", "r", "r", "r")
)
```
#### Cell Code
```{webr-r}
#| context: interactive
#| label: part-b-chunk-14
library(dplyr)
library(ggplot2)
library(purrr)
library(tibble)
library(tidyr)
set.seed(2025)
# Realistic age distribution and MAR missingness:
# performance is more likely to be missing for older participants
age_raw <- round(rnorm(25, mean = 38, sd = 12))
age_vals <- pmax(18L, pmin(65L, age_raw))
perf_full <- round(40 + 0.6 * age_vals + rnorm(25, 0, 10))
miss_prob <- plogis((age_vals - 42) / 10)
perf_obs <- ifelse(runif(25) < miss_prob * 0.55, NA_real_, perf_full)
study_data <- tibble(
participant = 1:25,
age = age_vals,
satisfaction = sample(c(3:7, NA), 25, replace = TRUE,
prob = c(0.15, 0.2, 0.25, 0.2, 0.15, 0.05)),
performance = perf_obs
)
study_data_display <- study_data %>%
mutate(across(c(satisfaction, performance), ~ if_else(is.na(.x), "NA", as.character(.x)))) %>%
slice(1:10)
smallsamplelab_apa_table(
"8.1",
"Simulated study dataset with missing values (rows 1 to 10)",
study_data_display,
note = "The full simulated dataset contains 25 participants; the excerpt is shown to illustrate the missing values before diagnosis.",
align = c("r", "r", "r", "r")
)
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: part-b-chunk-14
library(tidyverse)
set.seed(2025)
# Realistic age distribution and MAR missingness:
# performance is more likely to be missing for older participants
age_raw <- round(rnorm(25, mean = 38, sd = 12))
age_vals <- pmax(18L, pmin(65L, age_raw))
perf_full <- round(40 + 0.6 * age_vals + rnorm(25, 0, 10))
miss_prob <- plogis((age_vals - 42) / 10)
perf_obs <- ifelse(runif(25) < miss_prob * 0.55, NA_real_, perf_full)
study_data <- tibble(
participant = 1:25,
age = age_vals,
satisfaction = sample(c(3:7, NA), 25, replace = TRUE,
prob = c(0.15, 0.2, 0.25, 0.2, 0.15, 0.05)),
performance = perf_obs
)
study_data_display <- study_data %>%
mutate(across(c(satisfaction, performance), ~ if_else(is.na(.x), "NA", as.character(.x)))) %>%
slice(1:10)
smallsamplelab_apa_table(
"8.1",
"Simulated study dataset with missing values (rows 1 to 10)",
study_data_display,
note = "The full simulated dataset contains 25 participants; the excerpt is shown to illustrate the missing values before diagnosis.",
align = c("r", "r", "r", "r")
)
```
::::
### Testing the MCAR Assumption
Little's MCAR test evaluates whether missingness is consistent with the MCAR mechanism. The test compares observed means across missing-data patterns. A large p-value suggests MCAR is plausible, whereas a small p-value indicates that missingness likely depends on observed data (i.e., not MCAR). With small samples (n < 50), Little's MCAR test has low power to detect meaningful departures from MCAR. A non-significant result does not confirm MCAR. Supplement it with visual inspection of missingness patterns, complete versus incomplete case comparisons, and substantive reasoning about why data might be missing [@graham2009].
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-b-data-collection-chapter-12-handling-missing-data-in-small-samples-cell-2"}
#### Rendered Output
```{r}
#| label: part-b-chunk-15-html
#| echo: false
# Little's MCAR test
mcar_output <- smalln_mcar_table(study_data)
smallsamplelab_apa_table(
"8.2",
"Little's MCAR test for the simulated study dataset",
mcar_output,
note = "A larger p-value is more consistent with MCAR, but with small samples this test is only one piece of evidence and does not confirm MCAR. If the naniar package is unavailable, the table records the number of observed missingness patterns and the formal test should be run before final reporting.",
align = c("r", "r", "r", "r")
)
```
```{r}
#| label: part-b-chunk-15a-html
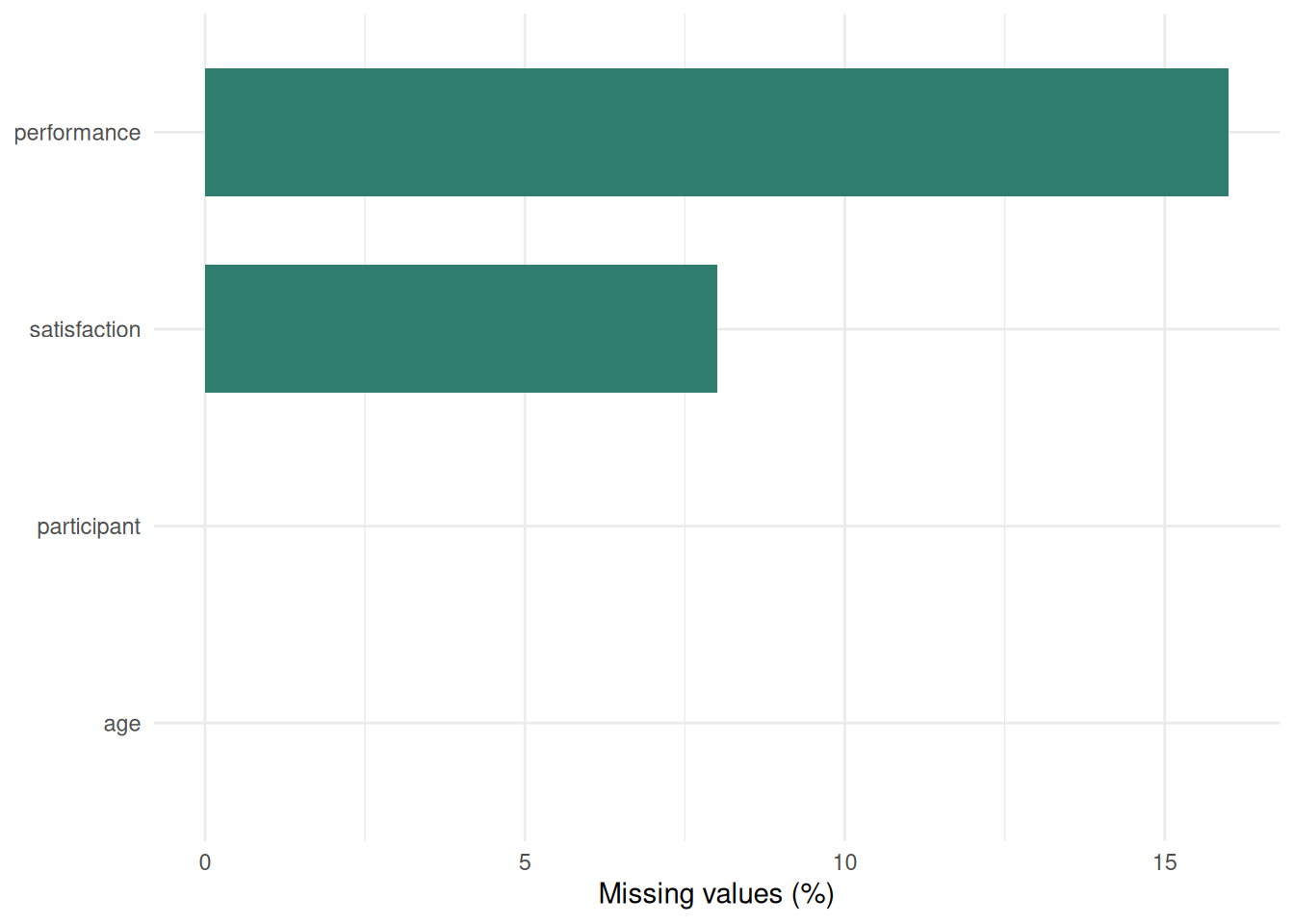
#| fig-cap: "Figure 8.1: Percentage of missing values per variable."
#| alt: "Percentage of missing values per variable."
#| echo: false
smalln_missing_var_plot(study_data)
```
#### Cell Code
```{webr-r}
#| context: interactive
#| label: part-b-chunk-15
#| fig-cap: "Figure 8.1: Percentage of missing values per variable."
# Little's MCAR test
mcar_output <- smalln_mcar_table(study_data)
smallsamplelab_apa_table(
"8.2",
"Little's MCAR test for the simulated study dataset",
mcar_output,
note = "A larger p-value is more consistent with MCAR, but with small samples this test is only one piece of evidence and does not confirm MCAR. If the naniar package is unavailable, the table records the number of observed missingness patterns and the formal test should be run before final reporting.",
align = c("r", "r", "r", "r")
)
smalln_missing_var_plot(study_data)
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: part-b-chunk-15
# Little's MCAR test
mcar_output <- smalln_mcar_table(study_data)
smallsamplelab_apa_table(
"8.2",
"Little's MCAR test for the simulated study dataset",
mcar_output,
note = "A larger p-value is more consistent with MCAR, but with small samples this test is only one piece of evidence and does not confirm MCAR. If the naniar package is unavailable, the table records the number of observed missingness patterns and the formal test should be run before final reporting.",
align = c("r", "r", "r", "r")
)
```
```{r}
#| label: part-b-chunk-15a
#| fig-cap: "Figure 8.1: Percentage of missing values per variable."
#| alt: "Percentage of missing values per variable."
smalln_missing_var_plot(study_data)
```
::::
Little's MCAR test assesses whether missing data patterns are completely random. Table 8.2 reports the test statistics, while Figure 8.1 shows the percentage of missing values for each variable. However, with small samples (n < 50), the test has low power and should be treated as one clue rather than a verdict. In practice, it should be supplemented with visual inspection of missingness patterns, comparisons between complete and incomplete cases, and domain knowledge about why participants might have skipped particular items or visits.
#### Observation-Level Missingness Pattern
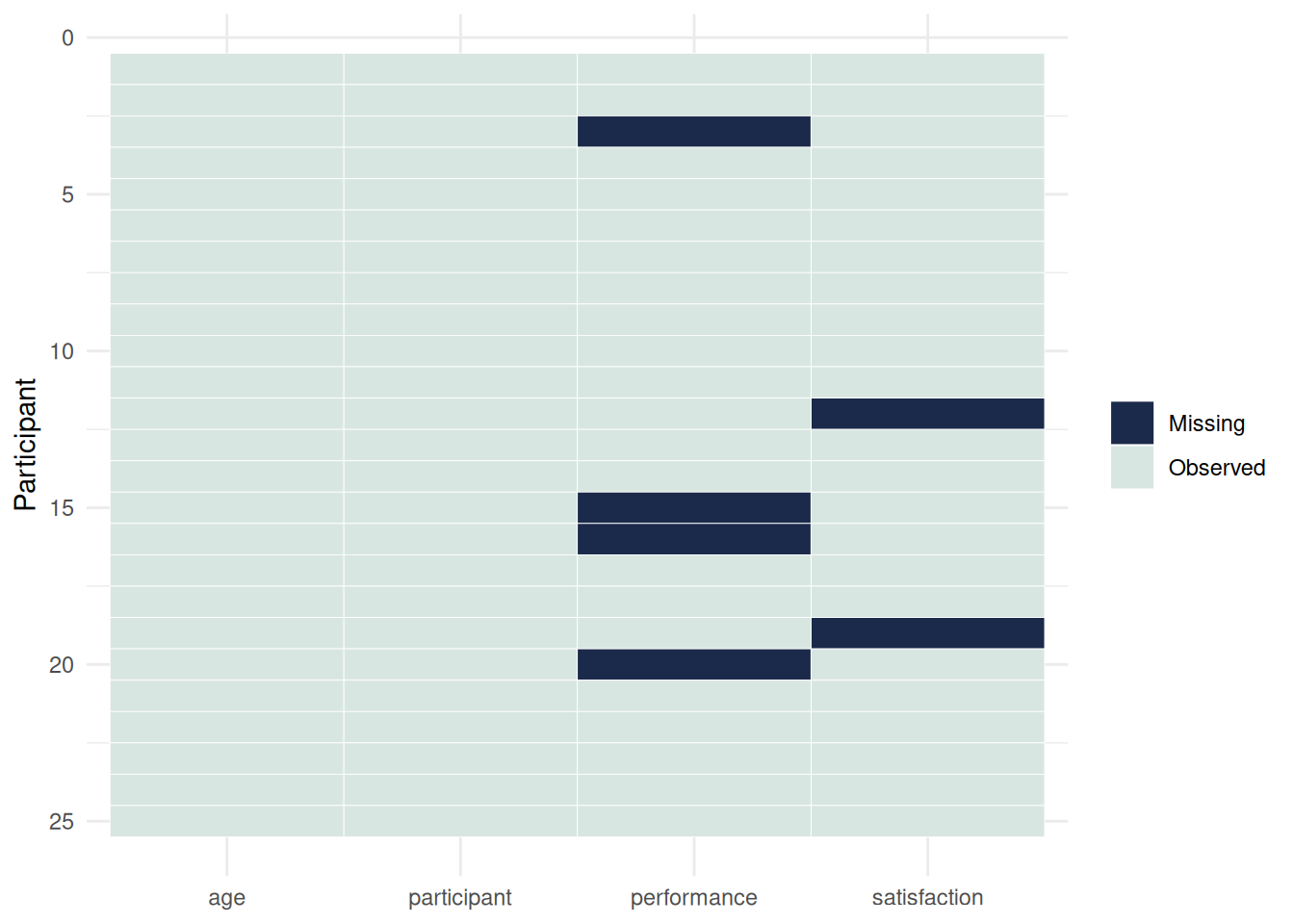
Figure 8.2 complements Figure 8.1 by showing whether missing values cluster within particular cases rather than only within particular variables.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-b-data-collection-chapter-12-handling-missing-data-in-small-samples-cell-2b"}
#### Rendered Output
```{r}
#| label: part-b-chunk-15b-html
#| fig-cap: "Figure 8.2: Observation-level missingness pattern."
#| alt: "Observation-level missingness pattern."
#| echo: false
smalln_missing_matrix_plot(study_data)
```
#### Cell Code
```{webr-r}
#| context: interactive
#| label: part-b-chunk-15b
#| fig-cap: "Figure 8.2: Observation-level missingness pattern."
smalln_missing_matrix_plot(study_data)
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: part-b-chunk-15b
#| fig-cap: "Figure 8.2: Observation-level missingness pattern."
#| alt: "Observation-level missingness pattern."
smalln_missing_matrix_plot(study_data)
```
::::
### Example: Summarising Missing Data
We continue working with the simulated dataset (`study_data`) created above. Tables 8.3 to 8.5 summarise how much data are missing and whether the incomplete cases differ from the complete cases on age.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-b-data-collection-chapter-12-handling-missing-data-in-small-samples-cell-3"}
#### Rendered Output
```{r}
#| label: part-b-chunk-16-html
#| echo: false
# Count missing values per variable
missing_summary <- study_data %>%
summarise(across(everything(), ~ sum(is.na(.)))) %>%
pivot_longer(everything(), names_to = "Variable", values_to = "Missing")
smallsamplelab_apa_table(
"8.3",
"Number of missing values by variable",
missing_summary,
align = c("l", "r")
)
# Proportion missing
prop_missing <- study_data %>%
summarise(across(everything(), ~ mean(is.na(.)))) %>%
pivot_longer(everything(), names_to = "Variable", values_to = "Proportion missing") %>%
mutate(`Proportion missing` = formatC(`Proportion missing`, format = "f", digits = 2))
smallsamplelab_apa_table(
"8.4",
"Proportion of missing values by variable",
prop_missing,
align = c("l", "r")
)
# Compare complete vs incomplete cases
study_data_with_complete <- study_data %>%
mutate(complete = complete.cases(study_data))
complete_vs_incomplete <- study_data_with_complete %>%
group_by(complete) %>%
summarise(mean_age = mean(age, na.rm = TRUE), n = n(), .groups = "drop") %>%
mutate(
complete = if_else(complete, "Complete", "Incomplete"),
mean_age = formatC(mean_age, format = "f", digits = 1)
) %>%
rename(Group = complete, `Mean age` = mean_age, n = n)
smallsamplelab_apa_table(
"8.5",
"Age comparison for complete and incomplete cases",
complete_vs_incomplete,
align = c("l", "r", "r")
)
```
#### Cell Code
```{webr-r}
#| context: interactive
#| label: part-b-chunk-16
# Count missing values per variable
missing_summary <- study_data %>%
summarise(across(everything(), ~ sum(is.na(.)))) %>%
pivot_longer(everything(), names_to = "Variable", values_to = "Missing")
smallsamplelab_apa_table(
"8.3",
"Number of missing values by variable",
missing_summary,
align = c("l", "r")
)
# Proportion missing
prop_missing <- study_data %>%
summarise(across(everything(), ~ mean(is.na(.)))) %>%
pivot_longer(everything(), names_to = "Variable", values_to = "Proportion missing") %>%
mutate(`Proportion missing` = formatC(`Proportion missing`, format = "f", digits = 2))
smallsamplelab_apa_table(
"8.4",
"Proportion of missing values by variable",
prop_missing,
align = c("l", "r")
)
# Compare complete vs incomplete cases
study_data_with_complete <- study_data %>%
mutate(complete = complete.cases(study_data))
complete_vs_incomplete <- study_data_with_complete %>%
group_by(complete) %>%
summarise(mean_age = mean(age, na.rm = TRUE), n = n(), .groups = "drop") %>%
mutate(
complete = if_else(complete, "Complete", "Incomplete"),
mean_age = formatC(mean_age, format = "f", digits = 1)
) %>%
rename(Group = complete, `Mean age` = mean_age, n = n)
smallsamplelab_apa_table(
"8.5",
"Age comparison for complete and incomplete cases",
complete_vs_incomplete,
align = c("l", "r", "r")
)
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: part-b-chunk-16
# Count missing values per variable
missing_summary <- study_data %>%
summarise(across(everything(), ~ sum(is.na(.)))) %>%
pivot_longer(everything(), names_to = "Variable", values_to = "Missing")
smallsamplelab_apa_table(
"8.3",
"Number of missing values by variable",
missing_summary,
align = c("l", "r")
)
# Proportion missing
prop_missing <- study_data %>%
summarise(across(everything(), ~ mean(is.na(.)))) %>%
pivot_longer(everything(), names_to = "Variable", values_to = "Proportion missing") %>%
mutate(`Proportion missing` = formatC(`Proportion missing`, format = "f", digits = 2))
smallsamplelab_apa_table(
"8.4",
"Proportion of missing values by variable",
prop_missing,
align = c("l", "r")
)
# Compare complete vs incomplete cases
study_data_with_complete <- study_data %>%
mutate(complete = complete.cases(study_data))
complete_vs_incomplete <- study_data_with_complete %>%
group_by(complete) %>%
summarise(mean_age = mean(age, na.rm = TRUE), n = n(), .groups = "drop") %>%
mutate(
complete = if_else(complete, "Complete", "Incomplete"),
mean_age = formatC(mean_age, format = "f", digits = 1)
) %>%
rename(Group = complete, `Mean age` = mean_age, n = n)
smallsamplelab_apa_table(
"8.5",
"Age comparison for complete and incomplete cases",
complete_vs_incomplete,
align = c("l", "r", "r")
)
```
::::
Interpretation: Tables 8.3 and 8.4 show that `performance` has the heaviest missingness burden, at about 20% of the sample. Table 8.5 then shows that the incomplete cases are younger on average than the complete cases, which makes MCAR less automatic and suggests that MAR or MNAR should remain plausible possibilities. With 20% missingness and n = 25, only 20 cases remain in a complete-case analysis, so power is reduced sharply.
### Complete-Case (Listwise Deletion) Analysis
The simplest approach is to analyse only cases with complete data on all variables of interest. This is valid if missingness is MCAR and the reduction in sample size is tolerable. However, it can introduce bias if missingness is MAR or MNAR, and it wastes information. Complete-case analysis is most defensible when missingness is minimal, MCAR is at least plausible, or the sample is so small that a more elaborate imputation model would be less credible than a transparent analysis of the observed cases.
::: {.callout-warning}
## Common Misconception: "Listwise Deletion Is Always Safe if Missingness Is Random"
**Myth**: "If I check for MCAR and the test is non-significant, listwise deletion is unbiased."
**Reality**: Even when missingness is **truly MCAR**, listwise deletion **loses power** and can introduce bias if you have multiple variables with independent missing patterns.
**Demonstration**:
```{r}
#| label: part-b-misconception-1
set.seed(2025)
# Generate complete data: n=50, correlation between x and y = 0.6
n <- 50
x <- rnorm(n, 50, 10)
y <- 0.6 * x + rnorm(n, 0, 8)
# True correlation (no missing data)
true_cor <- cor(x, y)
# Introduce MCAR missingness (20% on x, 20% on y, independently)
x_missing <- x
y_missing <- y
x_missing[sample(1:n, 10)] <- NA # 20% missing
y_missing[sample(1:n, 10)] <- NA # 20% missing
# Listwise deletion: only cases with both x and y
complete_cases <- complete.cases(x_missing, y_missing)
# Correlation with listwise deletion
listwise_cor <- cor(x_missing[complete_cases], y_missing[complete_cases])
listwise_demo <- tibble(
Metric = c(
"True correlation (complete data)",
"Complete cases retained",
"Correlation after listwise deletion",
"Cases lost",
"Standard error inflation"
),
Value = c(
formatC(true_cor, format = "f", digits = 3),
sprintf("%d / %d (%.1f%%)", sum(complete_cases), n, 100 * mean(complete_cases)),
formatC(listwise_cor, format = "f", digits = 3),
sprintf("%d (%.1f%%)", n - sum(complete_cases), 100 * (1 - mean(complete_cases))),
sprintf("%.2f×", sqrt(1 / sum(complete_cases)) / sqrt(1 / n))
)
)
smallsamplelab_apa_table(
"8.6",
"Consequences of listwise deletion under independent MCAR missingness",
listwise_demo,
note = "With 20% missing on x and 20% missing on y, independent MCAR retention is (1 - p1) x (1 - p2) = 0.8 x 0.8 = 0.64, so only about 64% of cases remain once both variables are required.",
align = c("l", "l")
)
```
**Why this matters:**
1. **Power loss**: With 20% missing on x and 20% on y independently, the retention rate is (1 - p_x) x (1 - p_y) = 0.8 x 0.8 = 0.64, so about 36% of cases are lost.
2. **Multiple variables compound**: With five variables each 15% missing, the retention rate is 0.85^5 = 0.44, so less than half the sample remains.
3. **Bias can still occur**: If missingness is MAR rather than MCAR, listwise deletion can bias estimates as well as reduce precision.
**Lesson**:
- **MCAR does NOT mean listwise deletion is optimal**—it can still waste information.
- Consider **multiple imputation** when missingness exceeds about 10%, the sample is large enough for the imputation model, and MAR is plausible.
- With small samples (n < 50), losing even 20% of cases can sharply reduce power.
**When listwise deletion is actually safe:**
- Missingness < 5% on any variable
- n is large enough that losing cases doesn't hurt power
- You've verified MCAR (not just MAR) AND documented the power loss
:::
### Mean Imputation (Not Recommended)
Mean imputation replaces missing values with the variable mean. This approach artificially reduces variance and distorts correlations. It is generally not recommended, especially with small samples where each imputed value has disproportionate impact. At most, it might be tolerated for trivial descriptive summaries in a much larger dataset, but it should not be treated as an inferential solution.
### Last Observation Carried Forward (LOCF)
In longitudinal studies, LOCF replaces missing follow-up values with the last observed value for that individual. This assumes no change after the last observation, which is often unrealistic. LOCF can therefore bias estimates and is not generally recommended. It is only marginally defensible when the assumption of no meaningful change is substantively plausible and no better alternative is available.
### Multiple Imputation (Caution with Small Samples)
Multiple imputation (MI) generates several plausible imputed datasets, analyses each separately, and pools results to account for imputation uncertainty. MI is a strong default for handling missing data in adequately sized samples when MAR is plausible [@rubin1987; @white2011; @vanbuuren2018]. If MNAR is suspected, MI alone does not solve the problem. The practical response is sensitivity analysis that varies the assumed departure from MAR [@sterne2009]. MI requires sufficient data to estimate imputation models reliably. With very small samples (n < 30) or extensive missingness (> 20%), imputation models may be under-identified and can yield unstable or implausible imputations. If MI is attempted in that setting, use predictive mean matching (`method = "pmm"`), limit the number of predictors in the imputation model, check convergence diagnostics carefully, and report complete-case results as a sensitivity check.
### Example: Multiple Imputation with mice (Caution)
We apply MI to the dataset with missing `satisfaction` and `performance` values. Given the small sample (n = 25) and 20% missingness, interpret results cautiously. Table 8.7 reports the pooled regression results rather than printing the raw imputation object.
:::: {.content-visible when-format="html"}
::::: {.panel-tabset group="part-b-data-collection-chapter-12-handling-missing-data-in-small-samples-cell-5"}
#### Rendered Output
```{r}
#| label: part-b-chunk-17-html
#| echo: false
# Multiple imputation requires the 'mice' package
if (requireNamespace("mice", quietly = TRUE)) {
library(mice)
# Remove 'complete' indicator variable before imputation
impute_data <- study_data %>% select(participant, age, satisfaction, performance)
# Perform multiple imputation (m = 5 imputations)
set.seed(2025)
imp <- mice(impute_data, m = 5, method = "pmm", printFlag = FALSE)
# Convergence diagnostic to run before reporting:
# plot(imp)
# Example analysis: regress performance on age and satisfaction
fit <- with(imp, lm(performance ~ age + satisfaction))
pooled <- pool(fit)
pooled_display <- summary(pooled) %>%
as_tibble() %>%
transmute(
Term = term,
Estimate = formatC(estimate, format = "f", digits = 3),
`Std. error` = formatC(std.error, format = "f", digits = 3),
Statistic = formatC(statistic, format = "f", digits = 2),
`p value` = formatC(p.value, format = "f", digits = 3)
)
smallsamplelab_apa_table(
"8.7",
"Pooled regression results from the illustrative multiple-imputation analysis",
pooled_display,
note = "Five predictive-mean-matching imputations were pooled using Rubin's rules. With only 25 cases and 20% missingness, these estimates should be treated as provisional.",
align = c("l", "r", "r", "r", "r")
)
} else {
smallsamplelab_apa_table(
"8.7",
"Multiple-imputation availability note",
tibble(Note = "Install the mice package to run multiple imputation. With very small samples, MI may be unstable; consider complete-case sensitivity analysis."),
align = "l"
)
}
```
#### Cell Code
```{webr-r}
#| context: interactive
#| label: part-b-chunk-17
# Multiple imputation requires the 'mice' package
if (requireNamespace("mice", quietly = TRUE)) {
library(mice)
# Remove 'complete' indicator variable before imputation
impute_data <- study_data %>% select(participant, age, satisfaction, performance)
# Perform multiple imputation (m = 5 imputations)
set.seed(2025)
imp <- mice(impute_data, m = 5, method = "pmm", printFlag = FALSE)
# Convergence diagnostic to run before reporting:
# plot(imp)
# Example analysis: regress performance on age and satisfaction
fit <- with(imp, lm(performance ~ age + satisfaction))
pooled <- pool(fit)
pooled_display <- summary(pooled) %>%
as_tibble() %>%
transmute(
Term = term,
Estimate = formatC(estimate, format = "f", digits = 3),
`Std. error` = formatC(std.error, format = "f", digits = 3),
Statistic = formatC(statistic, format = "f", digits = 2),
`p value` = formatC(p.value, format = "f", digits = 3)
)
smallsamplelab_apa_table(
"8.7",
"Pooled regression results from the illustrative multiple-imputation analysis",
pooled_display,
note = "Five predictive-mean-matching imputations were pooled using Rubin's rules. With only 25 cases and 20% missingness, these estimates should be treated as provisional.",
align = c("l", "r", "r", "r", "r")
)
} else {
smallsamplelab_apa_table(
"8.7",
"Multiple-imputation availability note",
tibble(Note = "Install the mice package to run multiple imputation. With very small samples, MI may be unstable; consider complete-case sensitivity analysis."),
align = "l"
)
}
```
:::::
::::
:::: {.content-visible unless-format="html"}
```{r}
#| label: part-b-chunk-17
# Multiple imputation requires the 'mice' package
if (requireNamespace("mice", quietly = TRUE)) {
library(mice)
# Remove 'complete' indicator variable before imputation
impute_data <- study_data %>% select(participant, age, satisfaction, performance)
# Perform multiple imputation (m = 5 imputations)
set.seed(2025)
imp <- mice(impute_data, m = 5, method = "pmm", printFlag = FALSE)
# Convergence diagnostic to run before reporting:
# plot(imp)
# Example analysis: regress performance on age and satisfaction
fit <- with(imp, lm(performance ~ age + satisfaction))
pooled <- pool(fit)
pooled_display <- summary(pooled) %>%
as_tibble() %>%
transmute(
Term = term,
Estimate = formatC(estimate, format = "f", digits = 3),
`Std. error` = formatC(std.error, format = "f", digits = 3),
Statistic = formatC(statistic, format = "f", digits = 2),
`p value` = formatC(p.value, format = "f", digits = 3)
)
smallsamplelab_apa_table(
"8.7",
"Pooled regression results from the illustrative multiple-imputation analysis",
pooled_display,
note = "Five predictive-mean-matching imputations were pooled using Rubin's rules. With only 25 cases and 20% missingness, these estimates should be treated as provisional.",
align = c("l", "r", "r", "r", "r")
)
} else {
smallsamplelab_apa_table(
"8.7",
"Multiple-imputation availability note",
tibble(Note = "Install the mice package to run multiple imputation. With very small samples, MI may be unstable; consider complete-case sensitivity analysis."),
align = "l"
)
}
```
::::
Interpretation: MI generates plausible values for missing data based on observed relationships. The pooled results combine estimates across imputations, with standard errors adjusted for imputation uncertainty. However, with n = 25 and 20% missingness, the imputation model is estimated from limited data, and results may be unstable. Before reporting the pooled estimates, inspect the standard `mice` trace plots with `plot(imp)` and increase `maxit` or simplify the imputation model if the chains drift rather than forming a stable fuzzy pattern. Compare MI results to complete-case analysis. If they differ substantially, report both and acknowledge uncertainty. Record the random seed, imputation method, number of imputations, and package versions whenever stochastic imputation code is adapted.
### Checking Convergence of Multiple Imputation
When using `mice`, always check whether the imputation algorithm has converged. Poor convergence means the imputed values may not be stable, especially with small samples or complex missing data patterns. This chapter keeps the emphasis on handling decisions rather than diagnostic graphics, so the detailed trace-plot and strip-plot workflow is taken up in Chapter 9: For the present chapter, the practical takeaway is that any MI analysis should be accompanied by those diagnostics before it is reported as credible.
### Sensitivity Analyses
When missingness is substantial or MNAR is suspected, conduct sensitivity analyses rather than presenting a single imputed answer as definitive. Compare complete-case results with imputed results, vary the assumptions about the missing-data mechanism where possible, and report how much the substantive conclusions change across those scenarios.
### Preventing Missing Data
The best approach to missing data is still prevention. Clear instruments, low respondent burden, follow-up for missed appointments or skipped questions, pilot testing of confusing procedures, and good rapport with participants all reduce the need for heroic statistical repair later.
### Key Takeaways
Missing-data work in small samples begins with description, not imputation. Researchers need to know how much is missing, where it is missing, and which missingness mechanisms are plausible before choosing a handling strategy. Complete-case analysis can be acceptable in narrowly defined situations but often wastes too much information, while multiple imputation is only as credible as the sample size, missingness level, and modelling assumptions allow. That is why prevention, diagnostics, and sensitivity analysis matter as much as the final pooled estimate.
### Self-Assessment Quiz
Test your understanding of missing-data decisions in Chapter 8.
```{r}
#| echo: false
#| results: asis
source(normalizePath(file.path(dirname(knitr::current_input(dir = TRUE)), "..", "R", "quiz_helpers.R"), mustWork = TRUE))
smallsamplelab_render_quiz(list(
list(
prompt = "What is the key distinction between MCAR and MAR?",
options = c("MCAR means no data are missing; MAR means some data are missing", "MCAR means missingness is unrelated to observed or unobserved data, whereas MAR allows missingness to depend on observed variables", "MCAR applies only to experiments; MAR applies only to surveys", "There is no practical difference between them"),
answer = 2L,
explanation = "The chapter defines MCAR as missingness unrelated to any variables and MAR as missingness related to observed variables but not the missing values themselves. This distinction matters because most modern missing-data methods assume MAR, not MCAR."
),
list(
prompt = "Why can listwise deletion still be a poor choice even when MCAR is plausible?",
options = c("It always creates impossible values", "It can throw away many cases and sharply reduce power, especially when missingness occurs on multiple variables", "It is only allowed in Bayesian analysis", "It automatically changes MAR data into MNAR data"),
answer = 2L,
explanation = "The chapter's misconception box shows that independent missingness across variables compounds quickly. Even under MCAR, listwise deletion can waste a large fraction of a small dataset and make estimates much less precise."
),
list(
prompt = "Why is mean imputation generally not recommended?",
options = c("It is too computationally expensive", "It artificially reduces variance and distorts associations between variables", "It requires a larger sample than multiple imputation", "It only works for binary outcomes"),
answer = 2L,
explanation = "Mean imputation replaces uncertainty with a constant value. The chapter warns that this shrinks variability and biases correlations, which is especially damaging when each observation matters."
),
list(
prompt = "When is multiple imputation most defensible in this chapter's guidance?",
options = c("When n is moderate, missingness is not too extensive, and MAR is plausible", "Whenever a dataset contains any missing value at all", "Only when data are MNAR", "Only when the outcome is binary"),
answer = 1L,
explanation = "The chapter describes multiple imputation as most appropriate when the sample is not extremely small, missingness is moderate, and the MAR assumption is plausible. It explicitly cautions that MI can be unstable with n < 30 or heavy missingness."
),
list(
prompt = "Why is last observation carried forward (LOCF) usually a weak solution to missing follow-up data?",
options = c("Because it assumes no meaningful change after the last observed value", "Because it can only be used with binary outcomes", "Because it always increases statistical power", "Because it requires Bayesian software"),
answer = 1L,
explanation = "The chapter warns that LOCF assumes the participant would have stayed unchanged after the last observed value. That assumption is often unrealistic, so LOCF can bias treatment effects or longitudinal trends."
),
list(
prompt = "What does Little's MCAR test evaluate?",
options = c("Whether missingness is consistent with a completely-random mechanism", "Whether multiple imputation has converged", "Whether the outcome is normally distributed", "Whether LOCF is acceptable"),
answer = 1L,
explanation = "Little's MCAR test compares observed means across missing-data patterns to assess whether the data are consistent with MCAR. The chapter also stresses that, with small samples, this test is only one clue rather than a final verdict."
),
list(
prompt = "Why are sensitivity analyses important for missing-data work?",
options = c("They prove which missingness mechanism is correct", "They show whether conclusions depend heavily on the assumptions used to handle missing data", "They eliminate the need to describe missingness patterns", "They allow you to ignore complete-case results"),
answer = 2L,
explanation = "Because the missingness mechanism is often uncertain, the chapter recommends comparing results under different reasonable assumptions. If results change materially, that uncertainty should be reported rather than hidden."
),
list(
prompt = "What is the best overall strategy for dealing with missing data in small-sample studies?",
options = c("Prevent as much missingness as possible through study design and follow-up", "Default to mean imputation because it is simple", "Always drop incomplete cases regardless of context", "Assume MCAR unless the sample is very large"),
answer = 1L,
explanation = "The chapter ends by stressing prevention: clear instruments, reduced burden, follow-up procedures, and pilot testing. No statistical fix can fully recover information that was never observed."
)
))
```